|
3/13/2023 0 Comments Seed data creator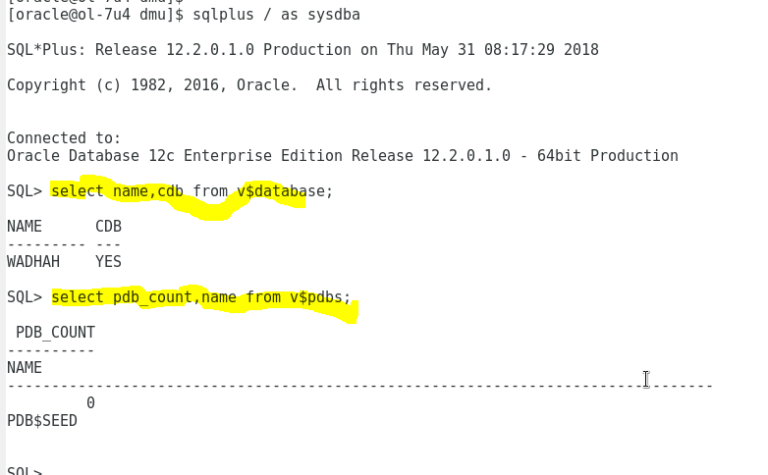
The third next(9) passes 9 into the generator as the result of the second yield and resumes the execution that reaches the end of the function, so done: true.…It reaches the second yield, that becomes the result of the generator call.next(4) passes 4 back to the generator as the result of the first yield, and resumes the execution. The result is returned to the outer code.next() starts the execution… It reaches the first yield. That argument becomes the result of yield. To do so, we should call generator.next(arg), with an argument. That’s because yield is a two-way street: it not only returns the result to the outside, but also can pass the value inside the generator. But in fact they are much more powerful and flexible. Until this moment, generators were similar to iterable objects, with a special syntax to generate values. It doesn’t use extra memory to store intermediate results. As a result, the `NDC1-01` files are in the `Data-VIS-20170203-1-room-light-off` folder.For (let i = start i <= end i++) yield i Ī generator composition is a natural way to insert a flow of one generator into another. However, the file was corrupted and hence, the acquisition was repeated during the batch `Data-VIS-20170203-1-room-light-off`. **Note:** The bundle `01` for the species `NDC1` was originally acquired during the batch `Data-VIS-20170111-2-room-light-off`. To permit possible registration between the two cameras, a chessboard pattern has been imaged and the acquired files are also contained in the folder `chessboard`. The HSI system was used to capture 256 wavelengths in this experiment and the exact wavelengths corresponding to the data provided are included in the file `wavelengths.csv`.īoth camera systems were fixed on a rigid frame for the duration of the experiments. Note: that there are 3 suffixes for each stem (`.hdr`, `.raw`, `.jpg`) Folder: The name of the folder containing the data (as described above where each folder contains a batch of images captured in a single imaging session). Bundle Number: Imaging Bundle Number (each bundle contains 48 kernels) every species has 2 bundles. Species Short Name: A shorthand of the species name. 
Species Full Name: The full species name (as used in filenames). 
The dark reference can be founds in each folder under the filename `black.hdr`/`black.raw`.Ī full index of the data for each species is provided in the `index.csv` file. For the dark reference, each folder contains an HSI image with the lens-cap covering the camera. To ensure stability, the halogen bulbs were switched on and allowed to reach constant operating temperature before the data were acquired in a dark room to minimise any other sources of illumination variance.įor the purposes of calibration each HSI image contains in the scene a 100% reflective spectralon tile which is a highly reflective Lambertian scatter. Two halogen bulbs were used for illumination and these were accurately positioned to provide balanced lighting across the scene. For instance the folder `Data-VIS-20170111-2-room-light-off` indicates that the data are in the VIS/NIR range, captured on the 11th of January 2017 and this was the second batch for that day with the room lights off. 
All the data from the same batch are contained in a dedicated folder. The data were captured in 9 batches across multiple days. For instance, the data for the `BC15` rice seed variety are contained in the following 6 files: 1 or 2), followed by the filename suffix. The filename convention used is the (short) species name followed by a dash, followed by the bundle number (i.e. `.hdr`: The HSI ENVI header file (More information on the ENVI format can be found at the () documentation. The following three files result from a single acquisition: This rice seed matrix was then positioned on a translational stage and imaged using the HSI and RGB cameras described above. For each imaging bundle, the 48 kernels were carefully positioned on a sheet of white paper and arranged in an `8圆` matrix. RGB - Fujifilm X-M1 with a 35mm/F2.0, ISO 400.įor each species, 96 kernels have been captured in two imaging bundles with 48 kernels in each bundle. Visible - Near Infrared (VIS/NIR) Hyperspectral Imaging Device System (~385nm - ~1000nm) consisting of a Specim V10E Imaging Spectrograph and Hamamatsu ORCA-05G CCD camera.Ģ. The dataset was collected in 2017 using the following two imaging systems:ġ. The dataset contains 90 rice seed species and 96 kernels per species resulting in 8,640 rice seed kernels in total. # RGB and VIS/NIR Hyperspectral Imaging Data for 90 Rice Seed Varieties
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
